by Matthew Taliaferro, University of Colorado Anschutz Medical Campus, [This article first appeared in The Conversation, republished with permission]
Before 2020, when my friends and acquaintances asked me what I study as a molecular biologist, their eyes would inevitably glaze over as soon as I said “RNA.” Now, as the COVID-19 pandemic has shown the power and promise of this molecule to the world at large, their eyes widen.
Despite growing recognition of the importance of RNA, how these molecules get to where they need to be within cells remains largely a mystery.
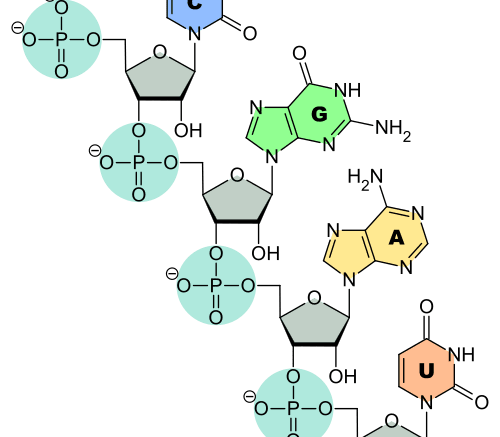
RNA is a chemical cousin of DNA. It plays many roles in the cell, but perhaps it’s most well-known as the relay messenger of genetic information. RNA takes a copy of the information in DNA from its storehouse in the nucleus to sites in the cell where this information is decoded to create the building blocks – proteins – that make cells what they are. This transport process is critical for animal development, and its dysfunction is linked to a variety of genetic diseases in people.
In some ways, cells are like cities, with proteins carrying out specific functions in the “districts” they occupy. Having the right components at the right time and place is essential.
For example, it makes little sense to put a high-security vault in the fashion district. Instead, it needs to be in the financial district, where there are tellers to fill it with currency. Similarly, proteins devoted to energy production for the cell are most functional not when they are confined to the nucleus but when they are in the cell’s power plant, the mitochondria, surrounded by the raw materials and accessories needed for their job.
The inside of a cell is much like a city.
So how do cells ensure the millions of proteins they contain are where they are supposed to be? One way is as simple as it sounds: transport them directly. However, every transport step costs energy. Dragging a heavy vault across town isn’t easy. An alternative strategy is to instead take the instructions for making the vault directly to the bank so it’s already in the correct location immediately after construction.
The instructions for making a given protein are contained within RNA. One way to ensure proteins are where they are supposed to be is to transport their RNA blueprint to where their specific functions are needed. But how does RNA get where it needs to be?
My research team focuses on this very question: What are the molecular mechanisms that control RNA transport? Our recently published research hints that some of the molecular language governing this process may be universal across all cell types.
The molecular language of RNA transport
For a handful of mRNAs – or RNA sequences coding for specific proteins – researchers have an idea about how they’re transported. They often contain a particular string of nucleotides, the chemical building blocks that make up RNA, that tell cells about their desired destination. These sequences of nucleotides, or what scientists refer to as RNA “ZIP codes,” are recognized by proteins that act like mail carriers and deliver the RNAs to where they are supposed to go.
My team and I set out to discover new ZIP codes that send RNAs to neurites, the precursors to the axons and dendrites on neurons that transmit and receive electrical signals. We reasoned that these ZIP codes must lie somewhere within the thousands of nucleotides that make up the RNAs in neurites. But how could we find our ZIP code needle in the RNA haystack?
Neurites are long, thin branches extending from the body of a neuron.
We started by breaking eight mouse neurite-localized RNAs into about 10,000 smaller chunks, each about 250 nucleotides long. We then appended each of these chunks to an unrelated firefly RNA that mouse cells are unlikely to recognize, and watched for chunks that caused the firefly RNA to be transported to neurites. To extend the mail analogy, we took 10,000 blank envelopes (firefly RNAs) and wrote a different ZIP code (pieces of neurite-localized RNA) on each one. By observing which envelopes were delivered to neurites, we were able to discover many new neurite ZIP codes.
We still didn’t know the identity of the protein that acted as the “mail carrier,” however. To figure this out, we purified RNAs containing the newly identified ZIP codes and observed what proteins were purified along with them. The idea was to catch the mail carrier in the act of transport while bound to its target RNA.
We found that one protein that regulates neurite production, named Unkempt, repeatedly appeared with ZIP code-containing RNAs. When we depleted cells of Unkempt, the ZIP codes were no longer able to direct RNA transport to neurites, implicating Unkempt as the “mail carrier” that delivered these RNAs.
Toward a universal language
With this work, we identified ZIP codes that sent RNAs to neurites (in our analogy, the bank). But where would an RNA containing one of these ZIP codes end up if it were in a cell that didn’t have neurites (a city that didn’t have a bank)?
To answer this, we looked at where RNAs were in a completely different cell type, epithelial cells that line the body’s organs. Interestingly, the same ZIP codes that sent RNAs to neurites sent them to the bottom of epithelial cells. This time we identified another mail carrier, a protein called LARP1, responsible for the transport of RNAs containing a particular ZIP code to both neurites and the bottom end of epithelial cells.
How could one ZIP code and mail carrier transport an RNA to two different locations in two very different cells? It turns out that both of these cell types contain structures called microtubules that are oriented in a very particular way. Microtubules can be thought of as cellular streets that serve as tracks to transport a variety of cargo in the cell. Importantly, these microtubules are polarized, meaning they have ingrained “plus” and “minus” ends. Cargo can therefore be transported in specific directions by targeting to one of these ends.
Microtubules are the roads proteins called kinesin use to transport materials from one cellular location to another.
In neurons, microtubules stretch through to and have their plus ends at the neurite tip. In epithelial cells, microtubules run from top to bottom, with their plus ends toward the bottom. Given that both of these locations are associated with the plus ends of microtubules, is that why we saw one ZIP code direct RNAs to both of these areas?
To test this, we inhibited the cell’s ability to transport cargo to the plus end of microtubules and monitored whether our ZIP code-containing RNAs were delivered. We found that these RNAs made it to neither the neurites in neurons nor to the bottom end of epithelial cells. This confirmed the role of microtubules in the transport of RNAs containing these particular ZIP codes. Rather than directing RNA to go to specific locations in the cell, these ZIP codes direct RNA to go to the plus ends of microtubules, wherever that might be in a given cell type.
We could compare this process to a mailing address. While the top line (“The Bank”) tells us the name of the building, it’s really the address and street name (“150 Maple Street”) that contains actionable information for the mail carrier. These RNA ZIP codes send RNAs to specific places along microtubule streets, not to specific structures in the cell. This allows for a more flexible yet uniform code, as not all cells share the same structures.
Moving mRNA into the clinic
Our research uncovers a new piece of how ZIP code sequences and proteins work together to get RNAs where they need to be. Our findings and methods can also be generalized to discover other new facets of the genetic ZIP code that direct RNAs to other locations in the cell.
Understanding how ZIP code sequences work can help researchers design RNAs that deliver their payload instructions to precise locations in the cell. Given the growing promise of RNA-based therapeutics, ranging from vaccines to cancer therapies, knowing how to make an RNA go from point A to point B is more important than ever.
Matthew Taliaferro, Assistant Professor of Biochemistry and Molecular Genetics, University of Colorado Anschutz Medical Campus
This article is republished from The Conversation under a Creative Commons license. Read the original article.